Identifying hit and lead molecules early on in the drug development process can make all the difference between a failure and a successful campaign. Typically, this is achieved through comprehensive screening campaigns with compound libraries screened directly against a drug target, or via more complex assays such as those involving cultured cells. In either case, extensive compound libraries must be assayed against the drug target via in silico, high throughput (HTS), or more focused screening techniques to identify hit molecules that bind or interact with the target in a desirable fashion.
However, assembling your custom compound library for a screening campaign can prove to be a substantial challenge, especially when the desired compound sets for a given screen need to be procured from multiple sources. This may leave you exposed to variability in lead time, compound format, shipping/import complications and price – not to mention the extra time spent communicating with multiple vendors. At Molport, our goal is to make the entire compound library assembly process easier by consolidating trustworthy vendors onto a single platform for compound sourcing and order management.
A powerful platform for research and procurement
In light of the current global supply chain crisis, screening compound procurement can seem like a difficult task. Molport can help you through these uncertain times, with our easy-to-use database that now includes 35 screening compound suppliers who have gone through our rigorous evaluation process prior to becoming our partners. This makes for a large diversity of choice when it comes to identifying the right screening sets for your research – with transparent pricing, lead time and stock availability at your fingertips.
Our database is comprised of over 10,000 unique natural products and over 100,000 natural-like products that are available both commercially and accessible to download for your in silico docking research. This powerful platform for screening compound research and procurement has been adopted by leading academic institutions, pharmaceutical, and biotech companies worldwide, and has enabled numerous success stories in recent years.
Let’s take a closer look at how our screening compound database has streamlined the drug discovery process for our valued customers.
Helping to identify drug candidates against SARS-CoV-2
The world came to a stop with the outbreak of COVID-19 in 2020, and prompted a race to develop therapeutic and prophylactic pharmaceuticals against the disease. Alongside the development of mRNA vaccines targeting the SARS-CoV-2 spike protein, a number of research groups were seeking to discover small molecules with the potential therapeutic characteristics against COVID-19. This was the case for several academic groups, who harnessed Molport’s screening compound database for in silico drug discovery.
In the pursuit of therapeutics for COVID-19, an Egyptian group used the Molport screening compound platform to search for potential inhibitors of the SARS-CoV-2 protease [1]. Initially, 5000 potential leads from the database were discovered using in silico docking studies. The lead compounds were later assessed by molecular dynamics and molecular mechanics binding calculations. Combined, the two calculations unearthed 9 natural-like small molecules from the Molport screening compound database, all showing high potency and binding affinity, along with suitable pharmaceutical characteristics.
Another noteworthy study, led by a research group in India [2], also took advantage of Molport’s screening compound database for computational biology and molecular modelling drug discovery approaches to identify lead molecules against the SARS-CoV-2 protease. Using our natural compound database, a structure-based virtual screening identified 341 lead compounds. Following drug likeness filtering, autodocking, ADMET and molecular mechanics tests, two compounds were identified that displayed protein–ligand complex stability in simulated fluid conditions.
These studies demonstrate the power of exploring Molport’s database of natural and natural-like products for in silico research to identify potential therapeutic agents. Once identified, it’s easy to use our natural compound platform to order your hit and lead compounds for in vitro research.
Discovering novel AChE inhibitors for Alzheimer’s
In recent years, the inhibition of acetylcholinesterase (AChE) has become a promising therapeutic strategy for treating the symptoms of Alzheimer’s disease (AD). AChE inhibitors have been found to reduce AD symptoms by increasing acetylcholine concentrations in the brain, which in turn improves patient memory and cognitive function.
One Germany-based research group set out to identify small molecules which could act as novel AChE inhibitors for the treatment of AD, and harnessed Molport’s natural compound database to do so [3]. In the first leg of the study, docking-based computational screening was carried out using Molport’s database to search for synthetic analogs of physostigmine and donepezil, two highly potent commercially available AChE inhibitors. 11 lead compounds were identified and characterized in vitro for their inhibitory activity against AChE. From this study, several promising inhibitors were identified, the most potent of which showed a five-fold improved inhibitory concentration versus the popular commercial AChE inhibitor rivastigmine.
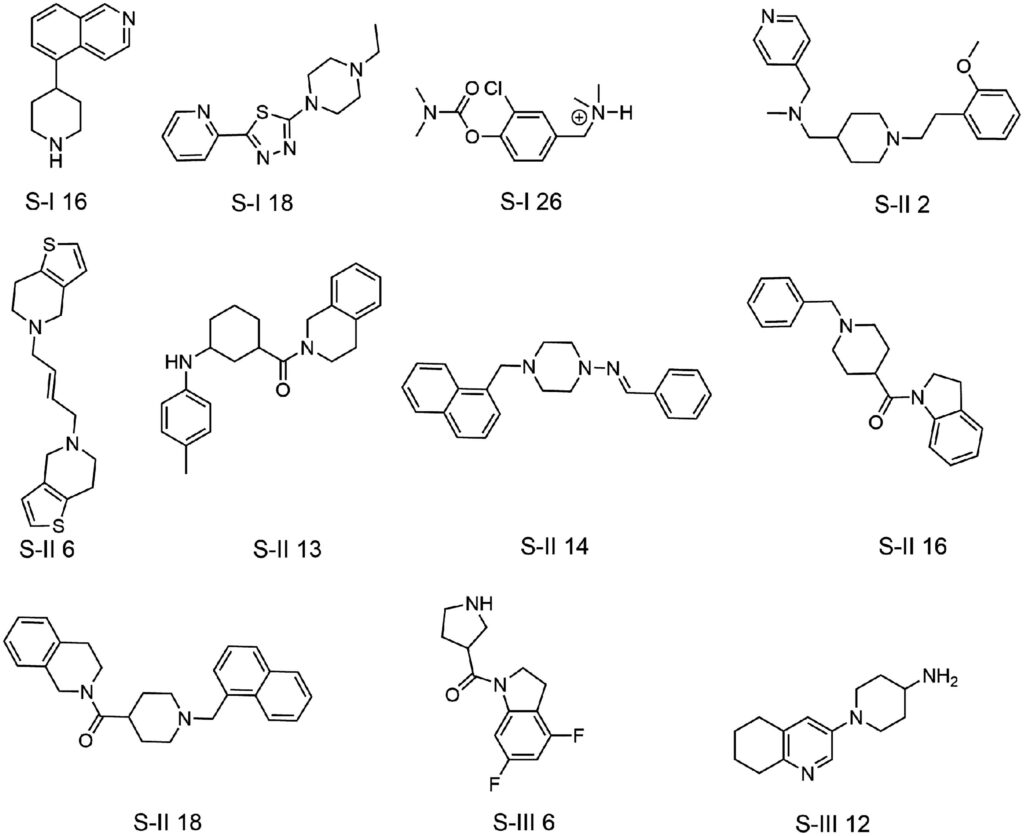
This study displays how Molport’s natural compound database can be harnessed for downstream in vitro characterization following an initial computational screen, and how the platform can be used to identify novel analogs of therapeutic drugs already on the market.
Identifying potent antimalarial hit molecules
Malaria is a significant public health burden in many tropical and subtropical regions, and accounts for approximately 400,000 deaths annually. With the prevalence of drug-resistant Plasmodium strains in the rise, there is an increasing need to explore novel antimalarial compounds with diverse modes of action. To this end, natural products have been widely exploited as a resourceful channel of biologically active compounds, demonstrated by antimalarial drugs such as quinine and artemisinin, which are derived from natural products.
One research group in Brazil set out to solve this problem, and looked once again towards natural screening compounds to identify those with potential antimalarial activity [4]. Molport’s natural compound database was employed for a quantitative structure–activity relationship (QSAR)-based virtual screening campaign, which returned 28 hits. Experimental validation confirmed potent and selective activity of two hit molecules identified in silico.
The results of this study support the use of Molport’s natural compound database for virtual screening campaigns to identify promising natural compound-based antimalarial molecules.
Glycolate oxidase inhibitors as a novel PH1 therapeutic
Primary hyperoxaluria type I (PH1) is a rare inherited renal disease that currently has few treatment options. Driven by a build-up of glyoxylate, crystals form in the renal tract as the disease progresses, resulting in kidney damage.
In a 2022 study [5], researchers from the University of Oxford, UK, used Molport compounds in a small molecule screening campaign to identify molecules against a novel therapeutic target – the enzyme glycolate oxidase. This enzyme is responsible for the breakdown of glyoxylate, and if targeted, could help to reduce glyoxylate buildup.
The researchers used Molport screening compounds in vitro to target the gating loop on glycolate oxidase, a promising allosteric binding site. Following crystallography-based fragment screening, they identified two hit compounds that were found to bind and inhibit glycolate oxidase via fluorescence-based activity assays and surface plasmon resonance.
This study demonstrates the promise of targeting an allosteric site to alter enzyme function. Indeed, it is the first report of allosteric inhibition of any hydroxy acid oxidase, and shows the potential of Molport gating loop target molecules for improving inhibitor selectivity for many other enzymes that could be implicated in disease.
Why not try our screening compound database today?
We have seen great success stories from our valued customers in recent years. These studies have come from various regions across the globe, and represent just a few of our operational domains. Many of these examples demonstrate the power of using our natural compound database for in silico screening campaigns, helping to identify potential therapeutic agents to address critical unmet medical needs.
Following a computational screening study, you can use our platform to easily order any hit and lead compounds for your lab-based support research. With over 4 million stock screening compounds, 410, 000 stock building blocks and counting, and the majority of customized quotes sent within 24 hours, it’s no wonder that many researchers are choosing a trusted and experienced partner like Molport.
References
- Ibrahim, M., Abdeljawaad, K., Abdelrahman, A., & Hegazy, M. F. (2021). Natural-like products as potential SARS-CoV-2 Mpro inhibitors: in-silico drug discovery. Journal of biomolecular structure & dynamics, 39(15), 5722–5734. https://doi.org/10.1080/07391102.2020.1790037
- Tripathi, N., Goel, B., Bhardwaj, N., Sahu, B., Kumar, H., & Jain, S. K. (2022). Virtual screening and molecular simulation study of natural products database for lead identification of novel coronavirus main protease inhibitors. Journal of biomolecular structure & dynamics, 40(8), 3655–3667. https://doi.org/10.1080/07391102.2020.1790037
- David, B., Schneider, P., Schäfer, P., Pietruszka, J., & Gohlke, H. (2021). Discovery of new acetylcholinesterase inhibitors for Alzheimer’s disease: virtual screening and in vitro characterisation. Journal of enzyme inhibition and medicinal chemistry, 36(1), 491–496. https://doi.org/10.1080/14756366.2021.1876685
- Ferreira, L. T., Borba, J., Moreira-Filho, J. T., Rimoldi, A., Andrade, C. H., & Costa, F. (2021). QSAR-Based Virtual Screening of Natural Products Database for Identification of Potent Antimalarial Hits. Biomolecules, 11(3), 459. https://doi.org/10.3390/biom11030459
- Mackinnon, S. R., Bezerra, G. A., Krojer, T., Szommer, T., von Delft, F., Brennan, P. E., & Yue, W. W. (2022). Novel Starting Points for Human Glycolate Oxidase Inhibitors, Revealed by Crystallography-Based Fragment Screening. Frontiers in chemistry, 10, 844598. https://doi.org/10.3389/fchem.2022.844598





